Although it’s a difficult topic to study, because the human species has experienced a number of population bottlenecks over time it’s assumed that rates of consanguinous (close kinship) marriage or partnership have been quite high at times in human history. Today, more than 10% of the world’s progeny is derived from consanguinous partnerships, especially within sub-Saharan Africa, the Middle East, and throughout parts of Asia. Children born of consanguinous parents have a 3.5% higher mortality rate, although it is difficult to tease out other effectors such as poverty [1].
Interestingly, however, there is some indication that consanguinous pairing may be protective for certain types of diseases as well.
One of the fascinating things that population geneticists have discovered is that smaller populations naturally tend to have less variability than larger ones (duh, right?). In human traits that are multifactorial, like autism for instance, some genetic variants can affect the expression of others. And in some cases, this can lead to a “covering up” of the effects of a genetic variant that, alone, would be harmful [2].

That is, until you marry someone outside of your village and– in the case of today– perhaps someone with whom you may share your last common ancestor thousands of years ago, you are unaware that one of your gene variants could be harmful. Intermixing decreases the likelihood that your partner shares the same genetic variants as you and also decreases the likelihood, thanks to sexual recombination, that your kids are going to inherit all those interacting variants together that help keep certain harmful diseases at bay.
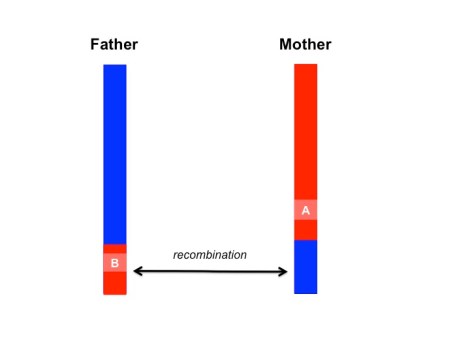
In the simplified scenario above, a segment of homologous chromosomes are sexually recombined from the mother and father. Paired variants, A and B, are unpaired, unmasking the expression of variant B.
While we’re not proposing that we start marrying our cousins again, understanding the links between large-scale population genetics and complex conditions like autism, heart disease, and cancer can help us to understand why they may be even more common. Intermixing across what were once isolated human populations helps to prevent rare recessive diseases, but it may also increase the likelihood that small-effect deleterious genetic variants can be unmasked.

That’s interesting, perhaps the best explanation for the reason why groups like the Amish experience a much lower (although not non-existent as has been posited) rate of autism.
Yes, it’s certainly a potential reason they tend to be plagued by recessive disorders but little autism and the like. Other groups to study would be some of the American gypsies, which are also known for certain recessive disorders.
Seriously?
Whereas autistic labelled symptomologies have emerged over the last forty years, intercommunal interbreeding has been rife for hundreds of generations and, in the “more developed” world it has been the norm for over a century.
Computers give us the power to handle vast multitudes of data, which, quite frankly, are so beyond our human interpretation that just because we can does not mean we should!
Genies are best left in their bottles……
Happy new year and the like. I’m pleased you continue to have your tea breaks – I drink far to much coffee…….
The level of travel and intermixing has exploded in the 20th century. But you already know my stance on autism prevalence and that it is not a new entity peculiar to the last century.
Still definitely drinking tea. Although I’ve switched from mugs and am enjoying a proper tea cup nowadays. 😊 👍🏻
But Emily, even in the last forty years the rate of increase, year on year has been phenomemal. That could never, ever be thought of as being a genetically sourced change. We are not drosophilla!
C/W the EXPLOSION in obesity and diabetes in US (and UK) since the 1980s. Also phenomenal. Also no way genetically derived.
I suspect that the rise in diagnosis rates, aside from being in part due to changes in how the label itself is defined, are affected by environmental factors. But I also suspect that heritable factors are involved in that vulnerability to environment. And small-effect genetic variants are actually the PERFECT partner to help explain vulnerability to environmental agencies. So the idea that common polymorphisms exist that affect autism risk would in fact fit perfectly with your focus on environmental influences.
Obesity and diabetes are pretty obvious and the US rates are in the exponential scales of increase. I’ll just mention Coeliac disease in passing, MS and such as well.
In an era when human health should be maximally perfect it blatantly not.
Whereas I deeply admire the/your ability to get right on in their to the core of the genetic sequences I question anyone, even the most powerful supercomputer’s ability to interpret them!
I mean, there is not even a clear definition of what actually designates “autism”.
Say, what d’you think of this:
https://www.sciencedaily.com/releases/2015/06/150601122445.htm
I agree. The lack of a good solid biological definition of autism is probably the most alarming issue in autism research, though few seem to recognize and acknowledge the emergency. To me at least, it’s the most important question. –Regarding the direct link between the immune system and CNS, I was pleased to read about that awhile ago. It certainly makes more sense. Although in researchers’ defense, the lymph nodes were apparently very easy to overlook.
For you my latest screed – mostly lifted from Dickie horton….
Pingback: Weekend Link Love – Edition 411 | Mark's Daily Apple·
Pingback: Weekend Link Love – Edition 411 | Health ,Beauty and Lifestyle·
Pingback: Weekend Link Love – Edition 411·
Pingback: Weekend Link Love – Edition 411 – Mark's Daily Apple·
Pingback: Time for another cup of Tea with Emily | greencentre·