Darwinism and the 20th century Modern Synthesis placed considerable emphasis on gradualism — the descent of species by slow, constant, often laborious modification. It’s an idea that’s been sold so well and so often that most media representations– and even many academic ones!– still portray evolution in this plodding, turtle-like manner.
But with the 1970s and Eldredge and Gould’s publication of their theory of Punctuated Equilibria (PE), evolutionary biologists began to view speciation as an infrequent occurrence, happening moreso during times of environmental upheaval but ultimately bookended by long periods of species stasis with only minor morphological fluctuations around a mean. In essence, the fossil record suggests that speciation is not like the slow, plodding tortoise– it is more like the hare, with long lazy naps in between short speedy dashes.

As part of my reading regimen during this extended time of COVID19 quarantine, I’ve been reading Ian Tattersall’s book, The Strange Case of the Rickety Cossack, which I recommend to anyone interested not only in human evolution but just evolutionary biology in general as he touches on a variety of major themes.

One of those ideas struck me in particular, explained by Tattersall in the following quote from the book:
Any feature you might want to follow in the fossil record is deeply embedded in a complex and autonomously functional organism that has to survive and compete in a multidimensional environment. Rarely will the fate of that organism depend entirely on the particular trait you’re interested in. What is more, the nature of heredity is such that every single gene plays a large variety of roles in the development of an organism, meaning that in most cases selection pressures cannot favor one particular aspect of a gene’s activity without also adventitiously affecting its other functions. And this integration of the genome brings us abruptly back to the fact that the target of natural selection is not any specific feature, but rather the survival and reproductive success of the entire organism. Natural selection– which is, after all, no more than differential reproduction– simply cannot single out particular traits to favor. It can only vote up or down on the whole package. You succeed or you fail in the reproductive stakes as the sum total of everything you are. … The bottom line is simple: individuals are such complex genomic and physical packages that within-population processes will rarely if ever be about optimization in specific features (pp. 107-108).
As many readers of this blog are probably already familiar, my background is not evolutionary biology or paleontology. Instead, I was originally trained in anatomy and neurobiology, with further emphasis on developmental biology and genetics. The vast majority of my work, to date, concerns the human condition.
As such, I’ve had the opportunity to study many rare genetic diseases that involve not only neurodevelopmental issues like autism and intellectual disability, but which are often accompanied by a host of other congenital anomalies [1]. Many of the genes associated with these conditions tend to be involved in developmental regulation (essentially, controlling the timing of developmental events). What’s more, most of the conditions I’ve studied are quite severe, in large part because they are often the result of some major loss-of-function mutation events, be they large copy number deletions that have wiped out entire genes or groups of neighboring genes, or even unlucky point mutations that result in premature stop codons and, essentially, truncated non-functional proteins.
As Dr. Tattersall states: “… every single gene plays a large variety of roles in the development of the organism.” And at least from my human studies, with their multiple congenital anomalies and variable intellectual and developmental problems, I find no statement could be truer.
However, I also have had the opportunity to study the feature landscape of many of these same genes, the structural components that ultimately lead to their unique expression and mutational patterns [2]. And so when Tattersall continues, saying, “… selection pressures cannot favor one particular aspect of a gene’s activity without also adventitiously affecting its other functions,” I must disagree.
There are some genes– close to 4,000– within the human genome that are considered “housekeeping genes” [3]. As you can probably guess by their name, these genes are involved in the daily upkeep and metabolism of cells. As such, because these functions are foundational to every cell within the body at every moment during its life, they are constitutively expressed. These genes have relatively short introns and contain comparatively few noncoding regulatory sequences that regulate aspects of their expression primarily because they are constantly expressed [2]. Complex patterns of regulatory elements are simply unnecessary for these genes.
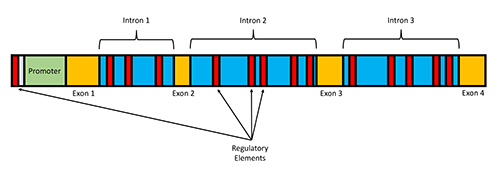
On the other hand, developmental genes– which are largely responsible for development of the physical form– are selectively expressed. As most biologists would say, they’re “tissue-specific.” As such, they contain larger introns and many many more noncoding regulatory elements within and around the genes that regulate their expressions in nuanced ways [2]. These regulatory elements effectively allow the genome to target gene expression according to cell type and even developmental time period. What’s more, these genes tend to produce many more transcript variations, which lead to different versions of the protein that can be used in different cell types under different instances and sometimes even for different purposes within the cell itself.
Therefore, mutations targeting these noncoding regulatory elements may actually allow evolution to fine tune its repertoire, targeting a limited number of cell types and time periods, while leaving others comparatively untouched– or at the very least, less impacted and therefore susceptible to further compensatory mutations [4].
What is perhaps even more interesting is that many of these regulatory sequences have been donated to the genome thanks to mobile element insertions and retention [5]. So punctuated influxes and expansions of mobile element families into organism’s genomes may actually speed along this whole process, providing material for the exaptation of more regulatory elements over time [6].
When a mutation occurs that causes a major loss-of-function within a developmental gene, it is typically– and very quickly– weeded out, either because the organism isn’t viable or reproduction is impaired in some way [reviewed in 7]. So, in this way, Tattersall is completely correct: evolution avoids major loss-of-function mutations. But mutations (including mobile element insertions and exaptations) targeting the more nuanced expression-related machinery of the genome may nevertheless offer a subtler backdoor option to evolution, allowing decoupling of fundamental processes and providing wiggle room within organismal development. In this way, evolution may– at least on occasion– hit a smaller target while not significantly impairing the organism’s ability to survive and reproduce.
